Search
Searching in WormBase ParaSite
Performing a search
The WormBase ParaSite search box is located in the top right on all pages:
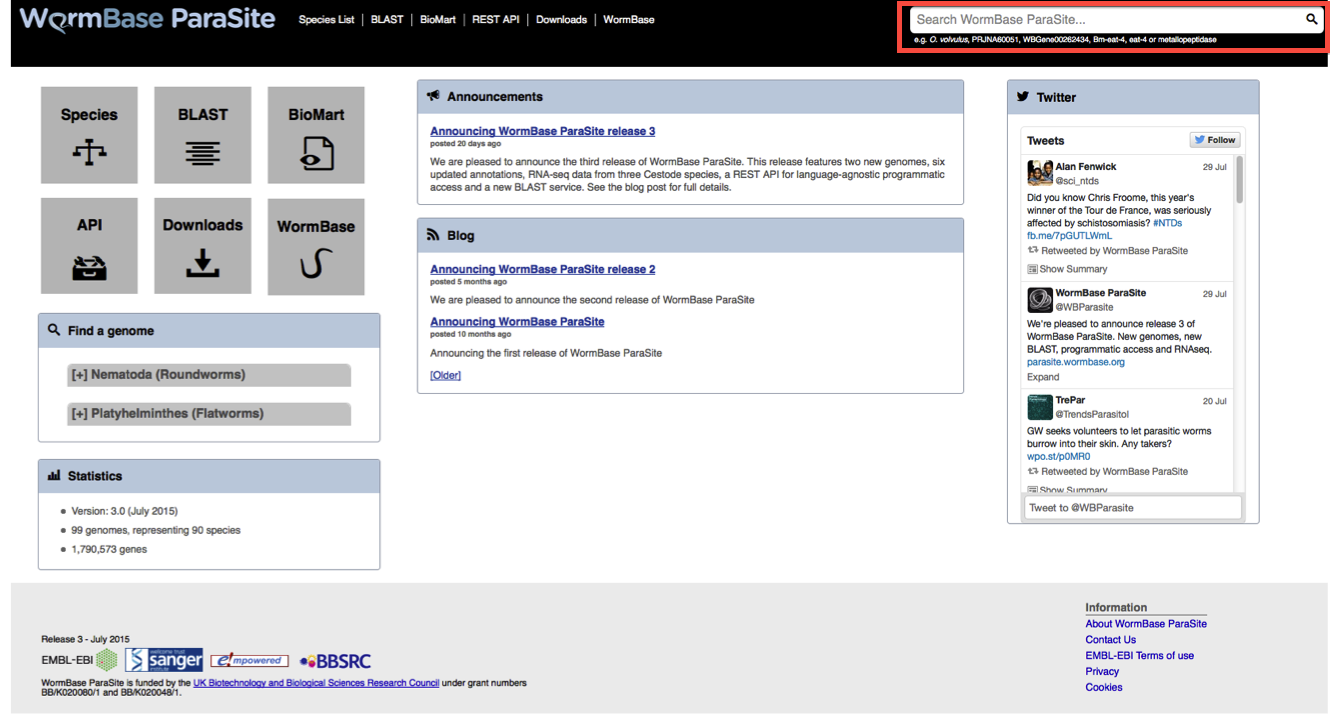
You can search on any field here, including species. If you start typing the autocomplete will give you a list of suggestions to choose from:
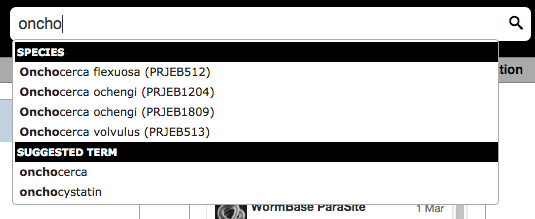
Search results
Gene search results are broken down into several fields:
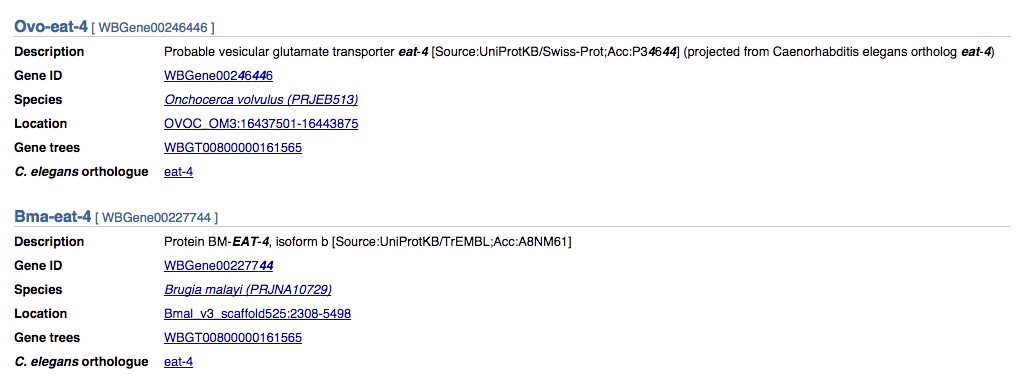
These are:
- Gene name
- Description of the gene and source information (e.g. projected from UniProt)
- Unique gene identifier
- Species name and genome project. Follow this link to take you to a page explaining a bit more about the species, details of how the genome was sequenced and what assembly/annotation has been performed.
- Location of the scaffold, contig or chromosome: follow this link to go to the genome browser
- Gene tree identifier. Follow this link to see the comparative genomics tree for the gene
- C. elegans ortholog (if present). Links back to the main WormBase website
Refining search results
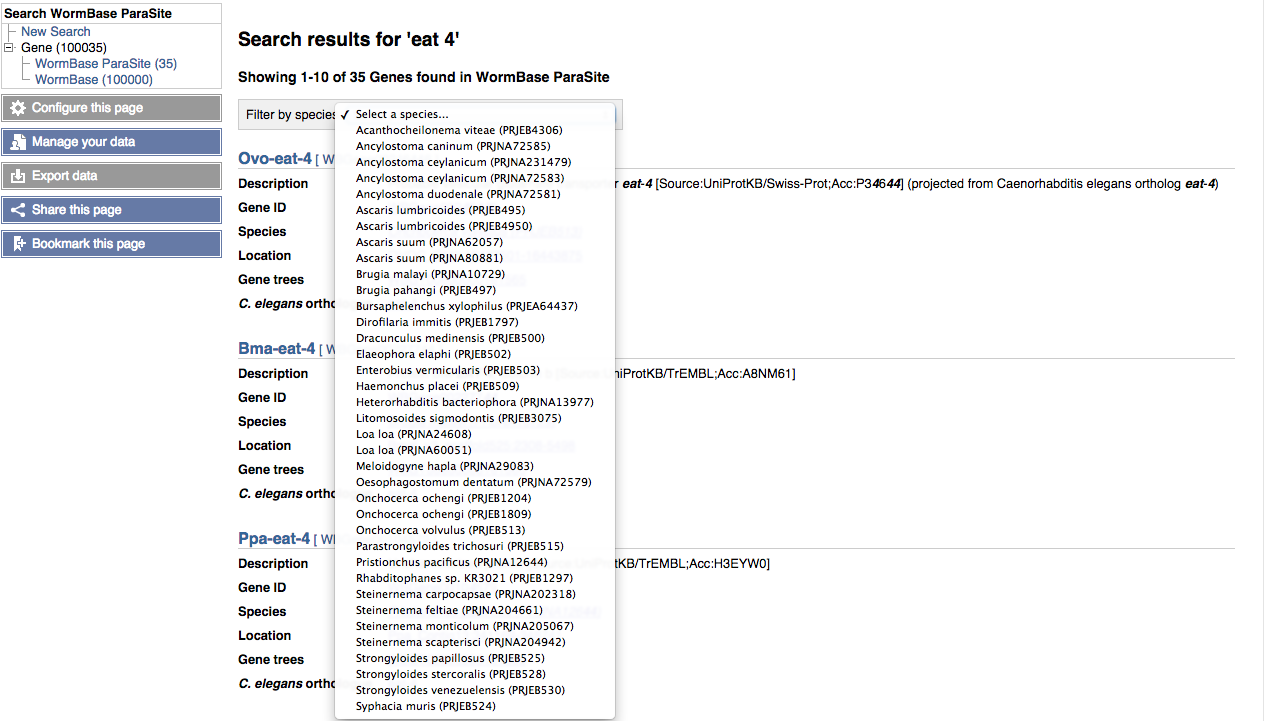
Search results are displayed as a list of either genes or species/projects. Gene search results can be filtered by species using the pulldown menu at the top of the list:
In addition to searching WormBase ParaSite, searches are also performed against the central WormBase site. The number of search results from each site are shown in the tree at the top left of the page:
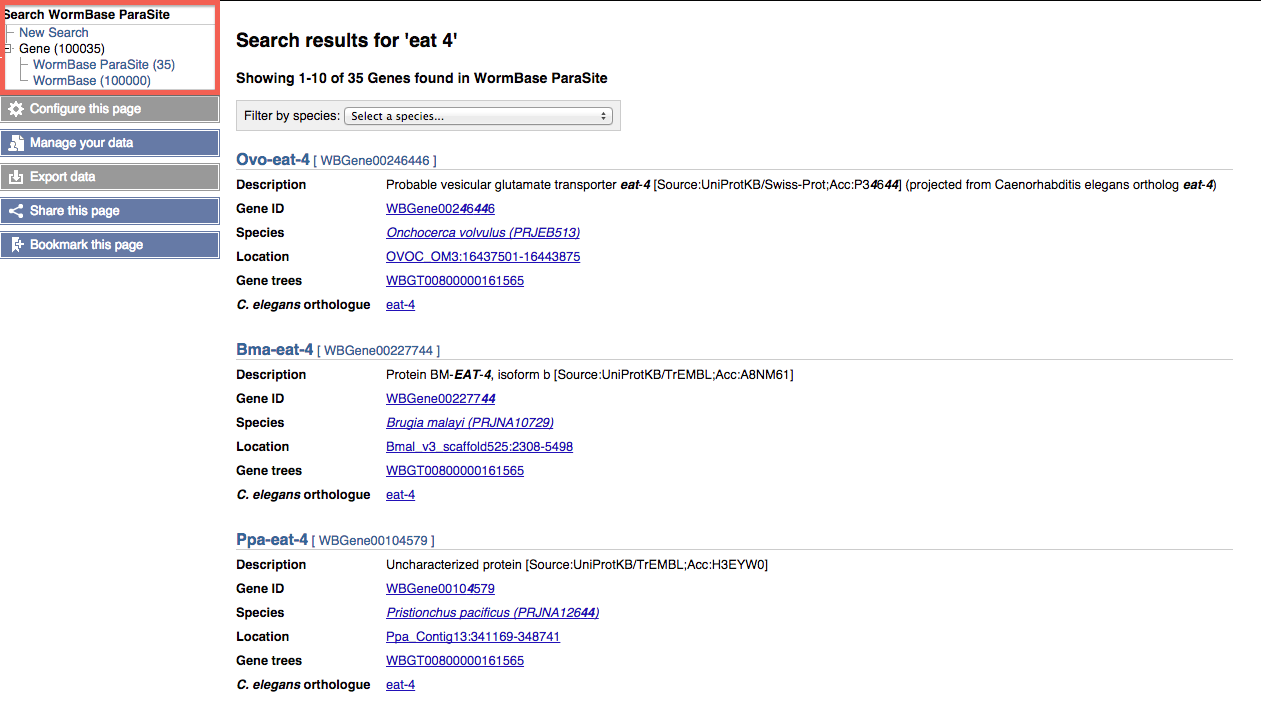
WormBase search results can be viewed by following the 'WormBase' link in the tree.